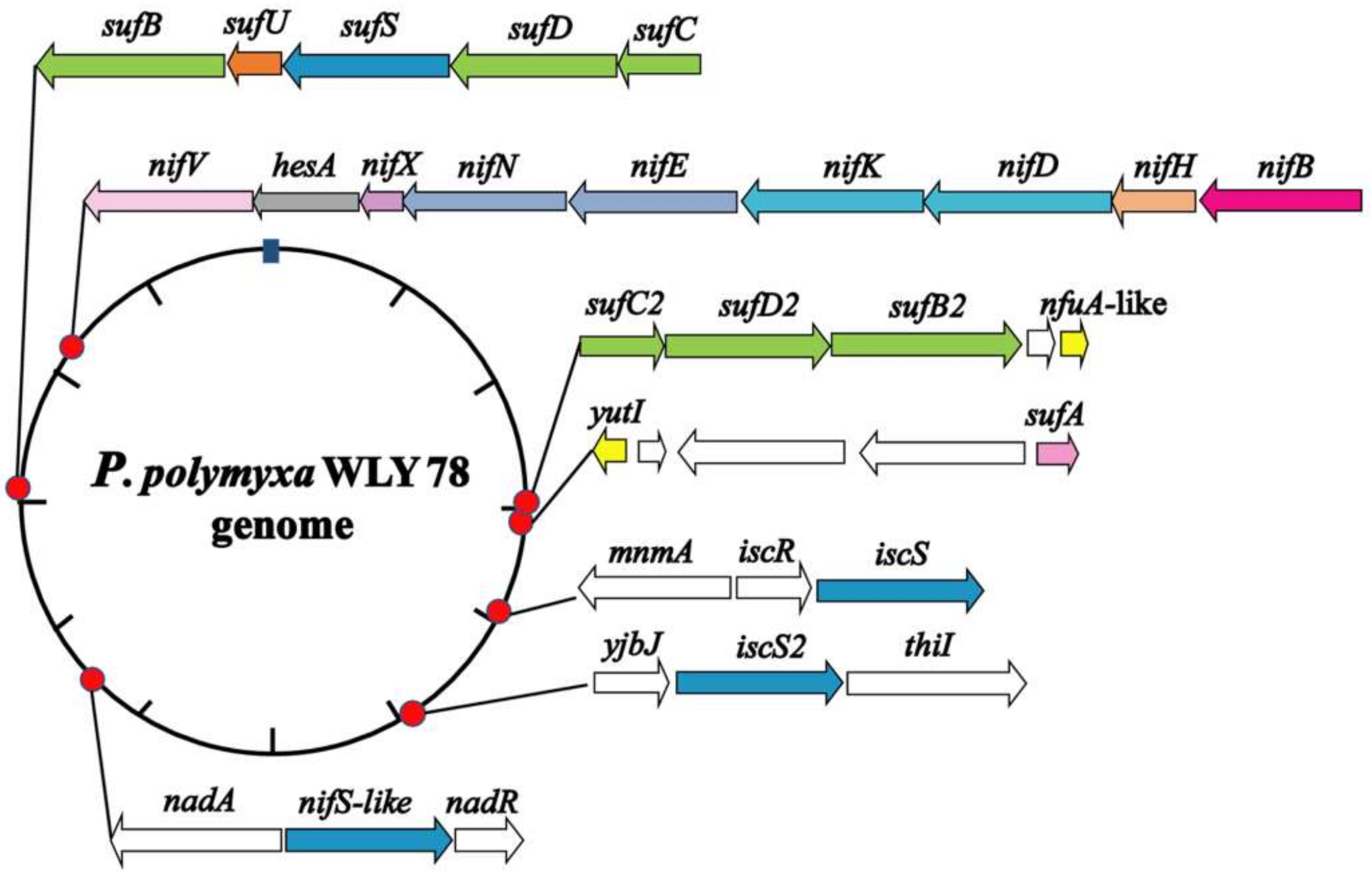
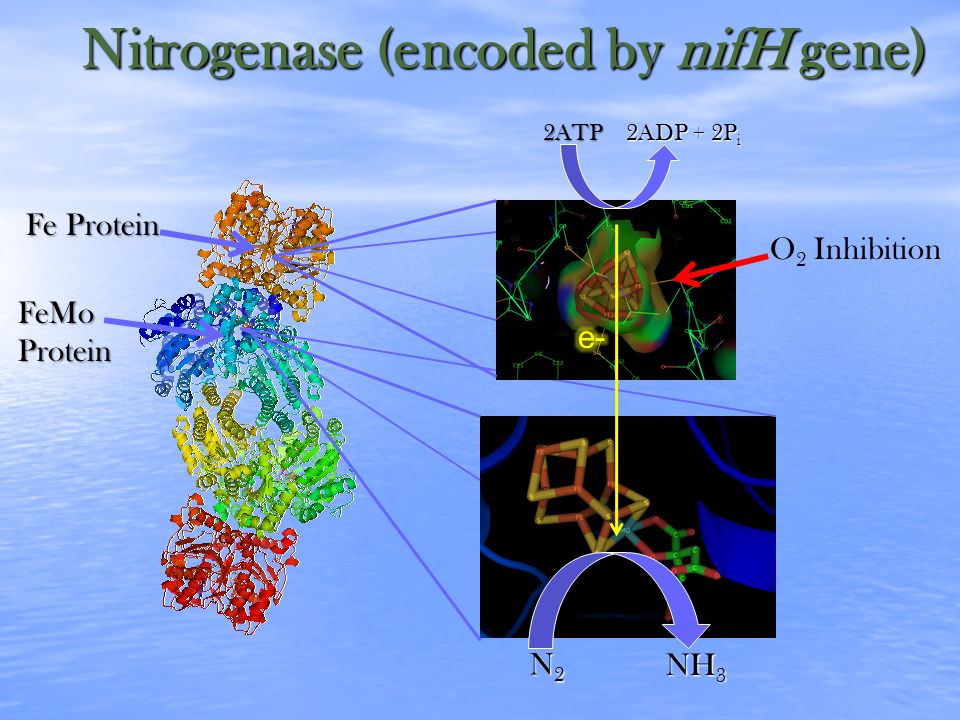
Abundance of nifH genes in urban, agricultural, and pristine prairie streams exposed to different levels of nitrogen loading | Semantic Scholar
![Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors](https://pub.mdpi-res.com/genes/genes-13-01428/article_deploy/html/images/genes-13-01428-g001.png?1660208416)
Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors
Molecular detection and quantification of nifH gene sequences in the rhizosphere of sorghum (Sorghum bicolor) sown with two leve
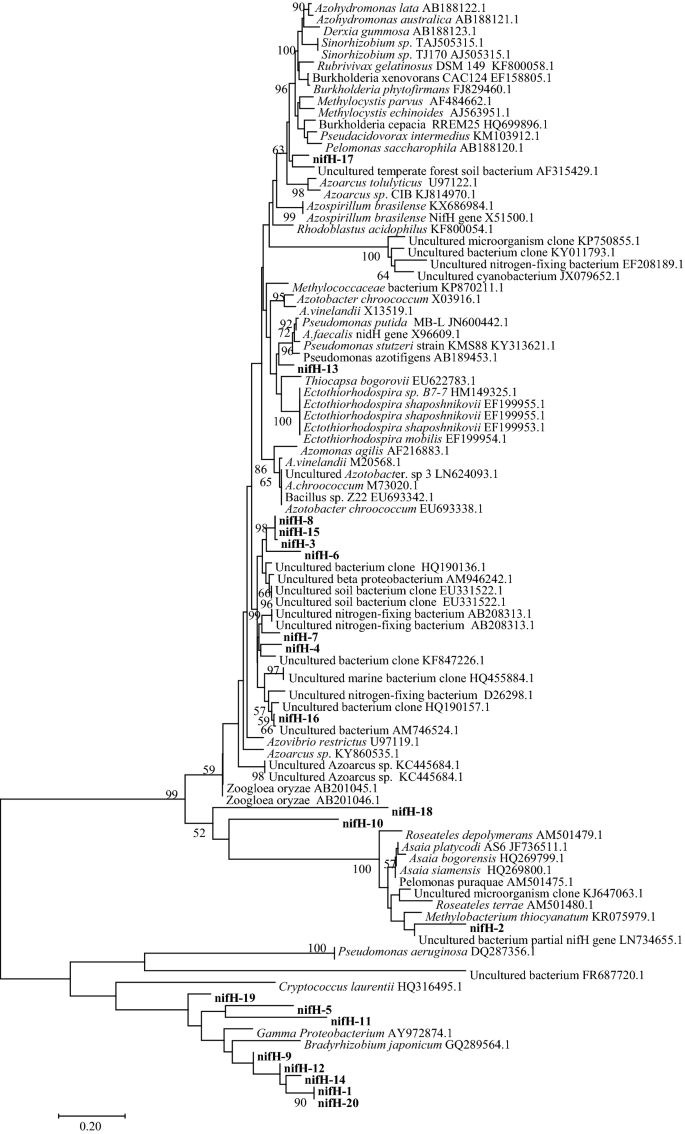
Functional diversity and abundance of nitrogen cycle-related genes in paddy soil | Applied Biological Chemistry | Full Text
Cloning, Prokaryotic Expression and Functional Characterization of NifH Gene from the Associative Nitrogen-Fixing Bacteria Klebsiella Variicola DX120E

Engineering Nitrogenases for Synthetic Nitrogen Fixation: From Pathway Engineering to Directed Evolution | BioDesign Research

The Flora Compositions of Nitrogen-Fixing Bacteria and the Differential Expression of nifH Gene in Pennisetum giganteum z.x.lin Roots
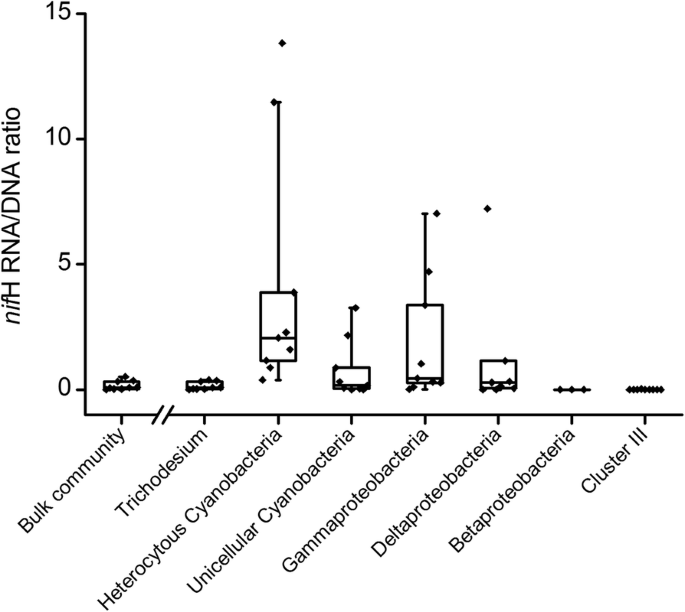
Analysis of nifH DNA and RNA reveals a disproportionate contribution to nitrogenase activities by rare plankton-associated diazotrophs | BMC Microbiology | Full Text

Frontiers | nifH Gene Sequencing Reveals the Effects of Successive Monoculture on the Soil Diazotrophic Microbial Community in Casuarina equisetifolia Plantations

Seasonal and long-term effects of nutrient additions and liming on the nifH gene in cerrado soils under native vegetation - ScienceDirect

Undervalued Pseudo-nifH Sequences in Public Databases Distort Metagenomic Insights into Biological Nitrogen Fixers | mSphere
![Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors](https://pub.mdpi-res.com/genes/genes-13-01428/article_deploy/html/images/genes-13-01428-g002.png?1660208413)
Genes | Free Full-Text | Genome-Wide Characterization of Nitrogenase Reductase (nifH) Genes in the Sweet Potato [Ipomoea batatas (L.) Lam] and Its Wild Ancestors

Formation of Nitrogenase NifDK Tetramers in the Mitochondria of Saccharomyces cerevisiae | ACS Synthetic Biology
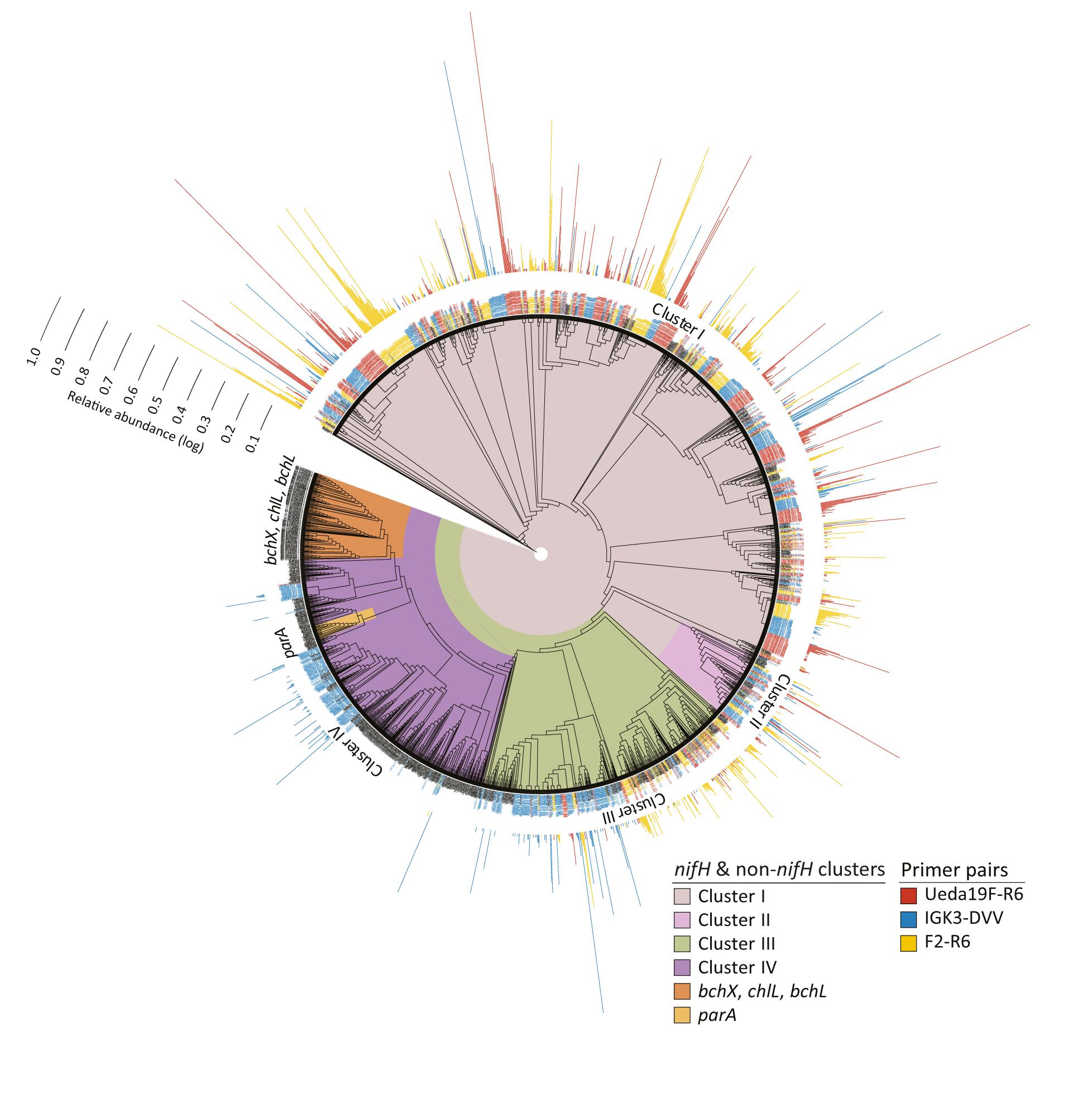
Frontiers | Evaluation of Primers Targeting the Diazotroph Functional Gene and Development of NifMAP – A Bioinformatics Pipeline for Analyzing nifH Amplicon Data

PCR amplification of nifH gene from soil and roots rice plant (~360 bp)... | Download Scientific Diagram