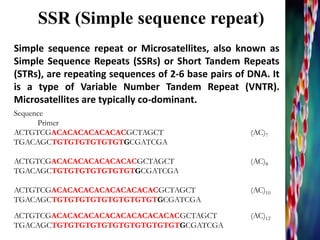
A general protocol for developing SSR markers with a SSR-enrichment step. | Download Scientific Diagram
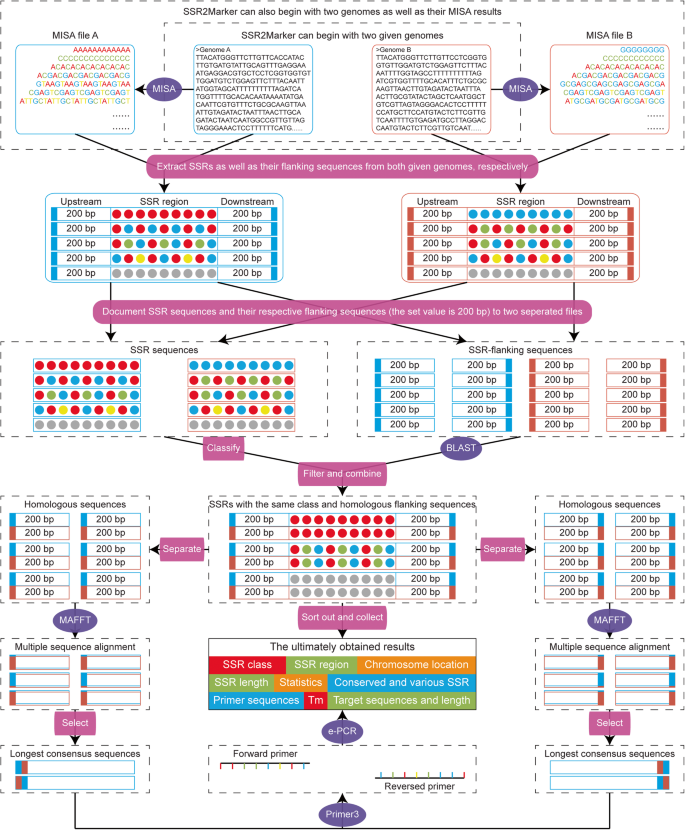
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text
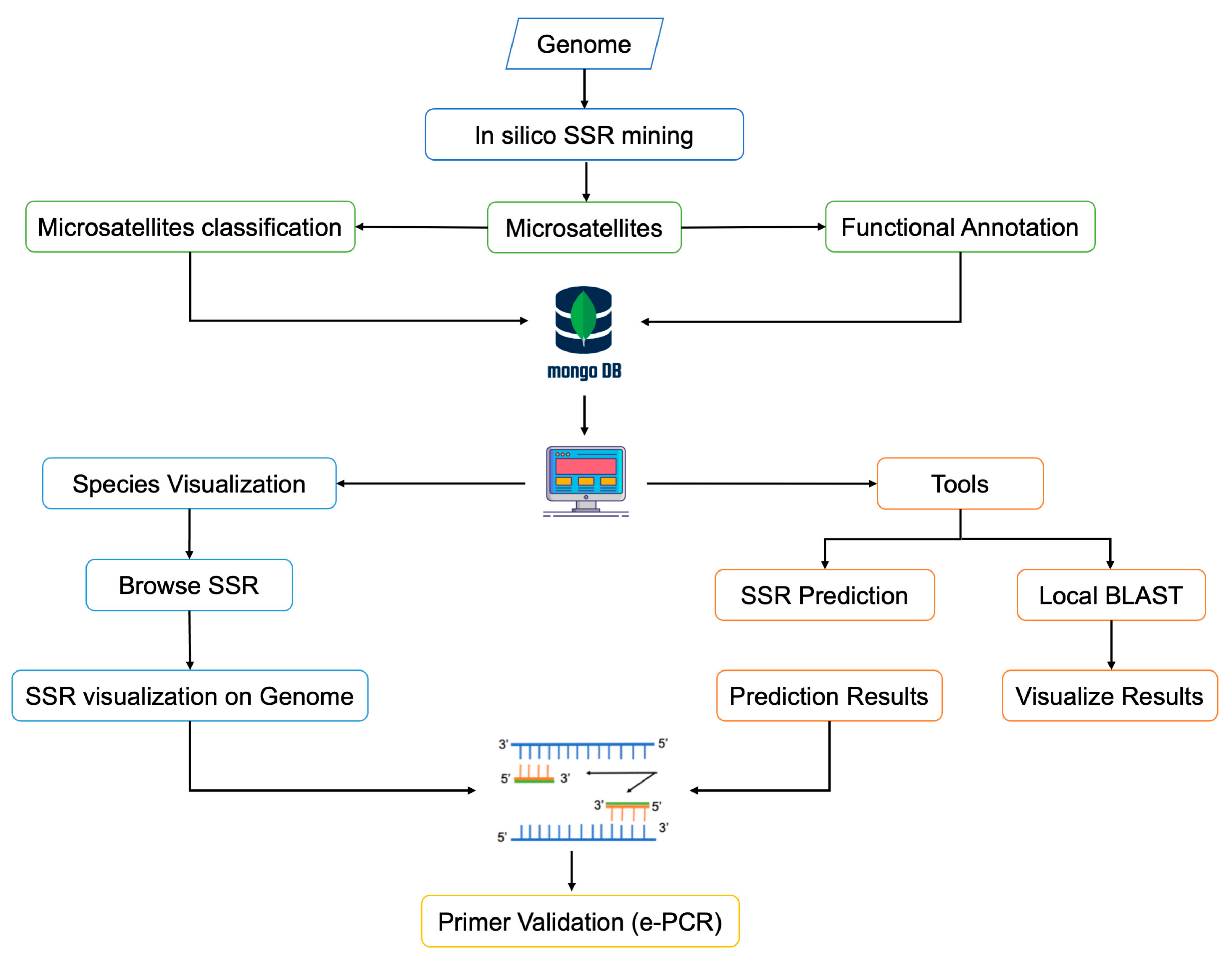
Genes | Free Full-Text | ranchSATdb: A Genome-Wide Simple Sequence Repeat ( SSR) Markers Database of Livestock Species for Mutant Germplasm Characterization and Improving Farm Animal Health
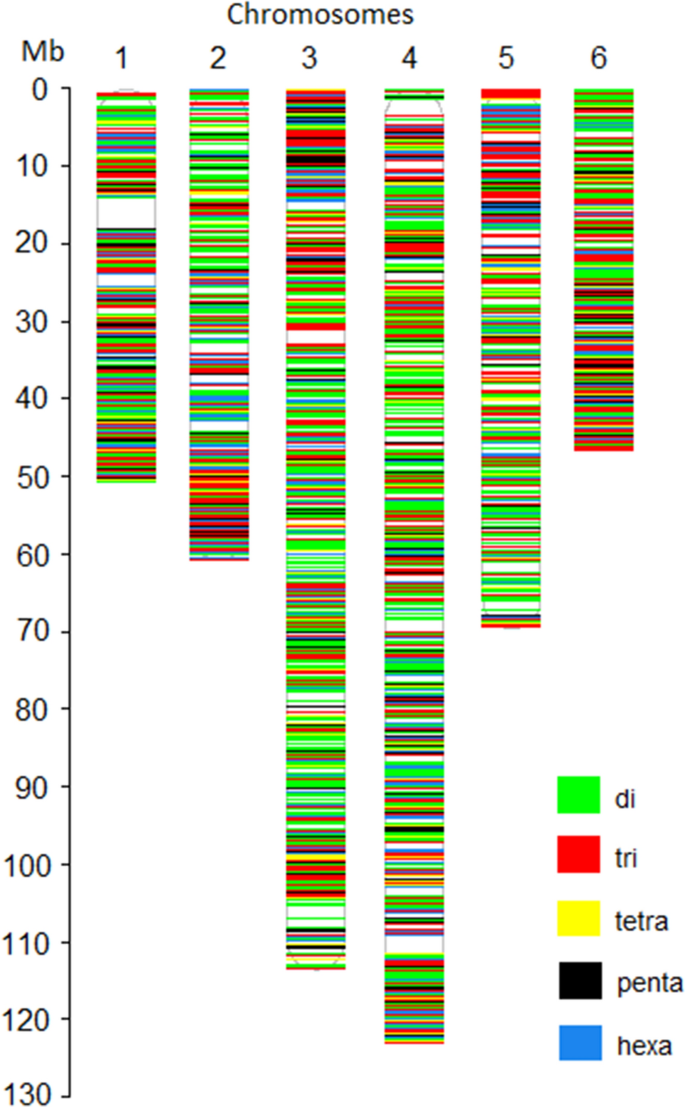
![Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh] Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]](https://www.frontiersin.org/files/Articles/243873/fpls-08-00377-HTML-r1/image_m/fpls-08-00377-g001.jpg)
Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]
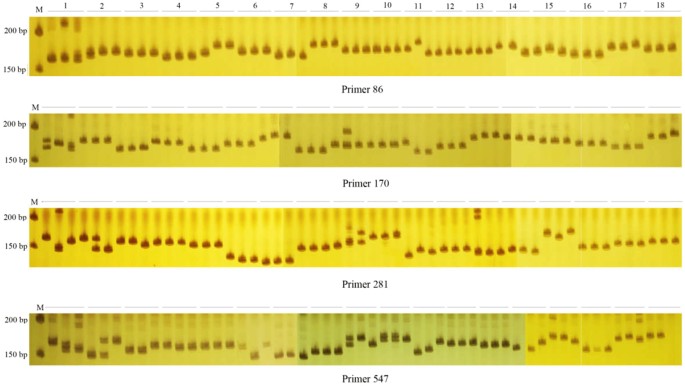

Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research | Scientific Reports

Diversity | Free Full-Text | SSR Markers for Trichoderma virens: Their Evaluation and Application to Identify and Quantify Root-Endophytic Strains
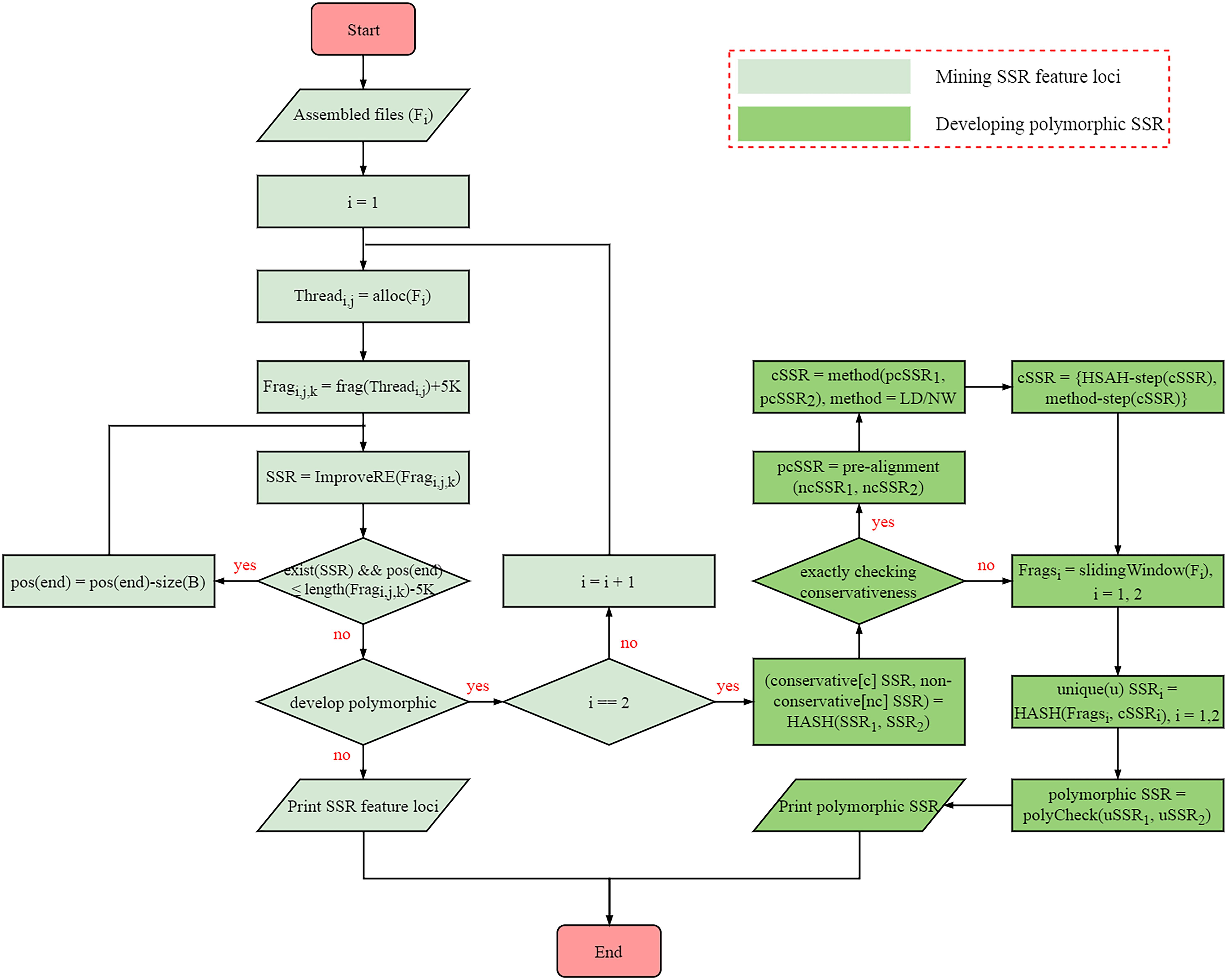
Frontiers | SSRMMD: A Rapid and Accurate Algorithm for Mining SSR Feature Loci and Candidate Polymorphic SSRs Based on Assembled Sequences

General flowchart of SSR marker discovery using different sequencing... | Download Scientific Diagram

Development of unigene-derived SSR markers in cowpea (Vigna unguiculata) and their transferability to other Vigna species
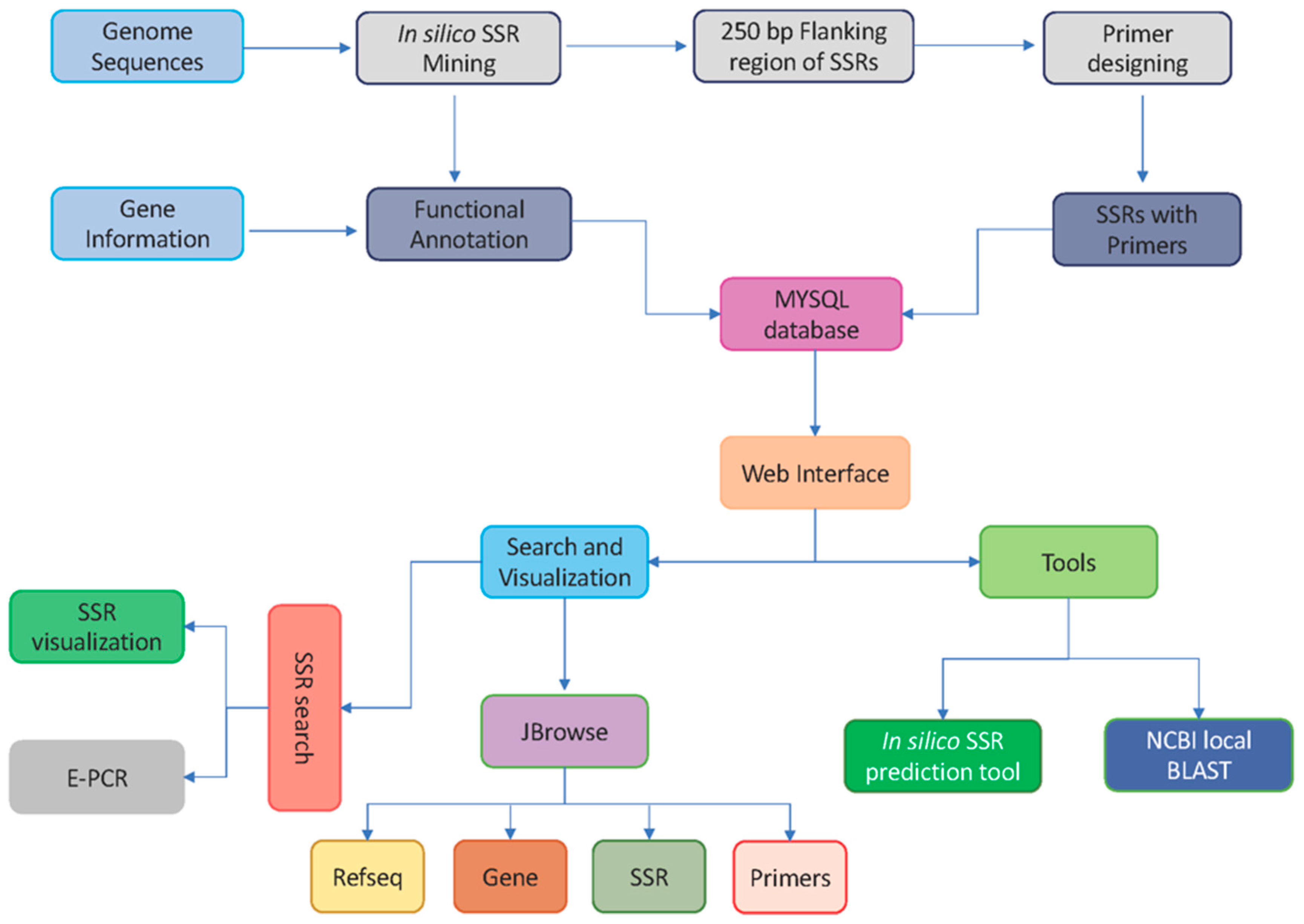
Genes | Free Full-Text | citSATdb: Genome-Wide Simple Sequence Repeat (SSR) Marker Database of Citrus Species for Germplasm Characterization and Crop Improvement

Characterization and Development of EST-SSR Markers from Transcriptome Sequences of Chrysanthemum (Chrysanthemum ×morifolium Ramat.) in: HortScience Volume 54 Issue 5 (2019)